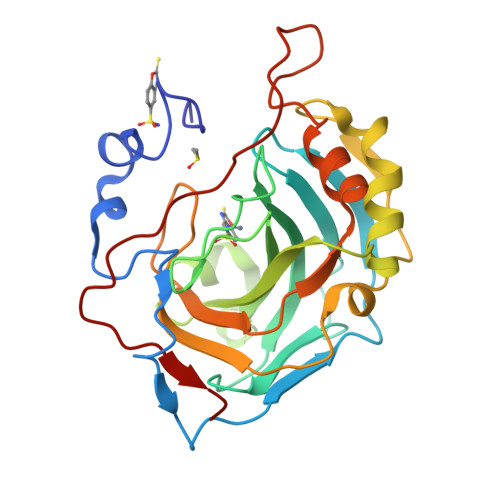
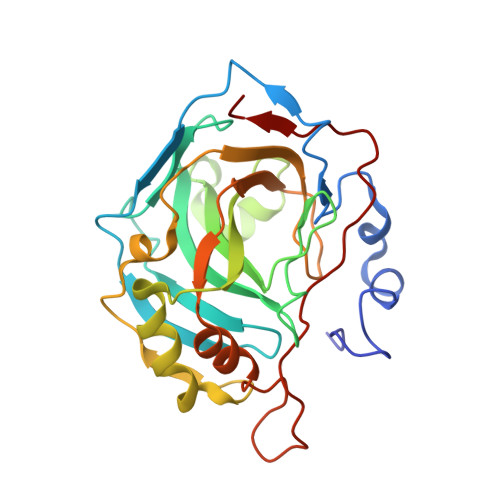
2-Mercaptobenzoxazoles: a class of carbonic anhydrase inhibitors with a novel binding mode to the enzyme active site.
Bozdag, M., Supuran, C.T., Esposito, D., Angeli, A., Carta, F., Monti, S.M., De Simone, G., Alterio, V.(2020) Chem Commun (Camb) 56: 8297-8300
- PubMed: 32573627
- DOI: https://doi.org/10.1039/d0cc02857f
- Primary Citation of Related Structures:
6YQT, 6YQU - PubMed Abstract:
2-Mercaptobenzoxazole is a widely used organic scaffold in medicinal chemistry. By means of kinetic and structural studies, we demonstrate that this molecule can effectively be used to inhibit hCAs showing a peculiar binding mode. The results reported here can pave the way for the development of selective CA inhibitors.
Organizational Affiliation:
Neurofarba Department, Section of Pharmaceutical and Nutriceutical Sciences, Università degli Studi di Firenze, Sesto Fiorentino, Florence, Italy.